Data science challenges in population genomics

Nearly a decade ago, the Human Genome Project successfully delivered a baseline definition of the DNA sequences in the entire human genome. Population genomics extends the scope of genomics research beyond baseline data to get a better understanding of gene variability at the level of individuals, populations, and continents.
Take India for example, where an ambitious program called IndiGen has been rolled out to map whole-genome sequences across different populations in the country. The first phase of the program, involving extensive computation analysis of the 1,029 sequenced genomes from India, identified 55,898,122 single-nucleotide variants in the India genome dataset, 32% of which were unique to the sequence samples from India. These findings are expected to provide the foundations for what will become an India-centric population-scale genomics initiative.
Population genomics opens up a range of region-specific opportunities such as identifying genes responsible for complex diseases, predicting and mitigating disease outbreaks, focusing on country-level drug development, usage, and dosing guidelines, and formulating precision public health strategies that deliver optimal value for the population.
As a result, several countries across the globe have launched their own initiatives for the large-scale comparison of DNA sequences in local populations.
The population genomics rush
Images Source: IQVIA
The International HapMap Project, launched in 2002 as a collaborative program of scientists from public and private organisations across six countries, is one of the earliest population-scale genomics programs.
A 2020 analysis of the global genomics landscape reported close to 190 global genomic initiatives, with the U.S. and Europe accounting for an overwhelming majority of these programs. Several countries have already launched large-scale sequencing programs such as All of Us (U.S.), Genomics England, Genome of Greece, DNA do Brasil, Turkish Genome Project, and the Saudi Human Genome Program, to name just a few.
Then there is the “1+ Million Genomes” initiative in the EU to create a cross-border network of national genome cohorts to unify population-scale data from several national initiatives.
There is a spectrum of objectives being collectively targeted by these projects including analysing normal and pathological genomic variation, improving infrastructure, and enabling personalised medicine.
As a result, population genomics data is exploding. An estimated 40 million human genomes have been sequenced as of 2020 with the number of analysed genomes expected to grow to 52 million by 2025. This exponential increase in population-scale data presents significant challenges, both in terms of crunching raw data at scale and in analysing and interpreting complex datasets.
The analytics challenge in population genomics
Genomic data volumes have been increasing exponentially over the past decade, thanks in part to the plummeting costs of next-generation sequencing technologies. Then there is the ever-expanding scope of health-related data, such as data from Electronic health records biomonitoring devices etc., that are becoming extremely valuable for population-scale research.
However, conventional integrative analysis techniques and computational methods that worked well with traditional genomics data are ill-equipped to deal with the unique data characteristics and overwhelming volumes of NGS and digital-era data. Data exploration and analysis already lag data generation by a significant order of magnitude – and that deficit will only be exacerbated as we transition from NGS to third-generation sequencing technologies.
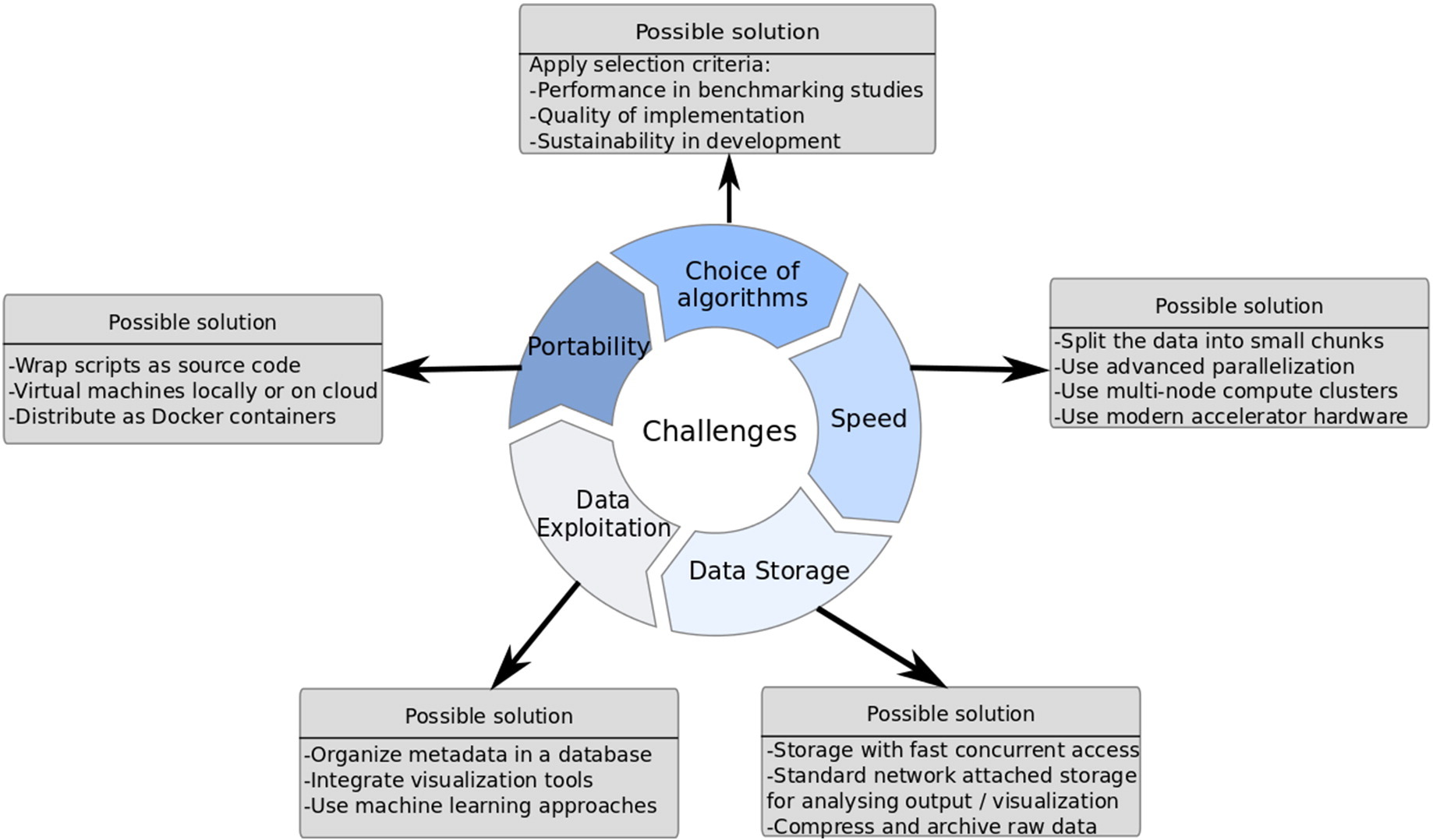
Image Source: ScienceDirect
Over the years, several de facto standards have emerged for processing genomics big data. But in spite of the significant progress that has been made in this context, the gap between data generation and data exploration continues to grow.
Most large institutions are already heavily invested in hardware/software infrastructure and in standardised workflows for genomic data analysis. A wholesale remapping of these investments to integrate agility, flexibility, and versatility features required for big data genomics is just plain impractical.
Integrating a variety of datasets from multiple external sources is a hallmark of modern genomics research and still represents a fundamental challenge for genomic analyses workflows.
The biggest challenge, however, is the demand for extremely specialized and scarce bioinformatics talent to build bespoke analytics pipelines for each research project. This significantly restricts the pace of progress in genomics research.
For data analysis to catch up with data acquisition, researchers need access to an easy-to-use powerful solution that spans the entire workflow – from raw data analysis to data exploration and insight.
The BioStrand “One Model” approach
At BioStrand, we offer an end-to-end, self-service SaaS platform that unifies all components of the genomics analysis and research workflow into one intuitive, comprehensive, and powerful solution. We designed the platform to address every pain point in the genomics research value chain.
For starters, it doesn't matter if you’re a seasoned bioinformatician or a budding geneticist. Our platform has a learning curve that’s as easy to master as Google search.
At BioStrand we believe that wrangling data is a tedious chore best left to technology. To that end, we have precomputed and indexed nearly 350 million sequences available across 11 public databases into one proprietary knowledge database that is continuously reviewed and updated. Ninety percent of population data from currently ongoing programs is soon expected to be publicly available, which means it will probably just be a click away.
In addition, you can add self-owned databases with just one click to combine them with publicly available datasets to accelerate time-to-insight. If it’s genomic data, we’ll make it computable.
With the BioStrand solution, you can use sequence or text to search through volumes of sequence data and instantly retrieve all pertinent information about alignments, similarities, and differences in sequences in a matter of seconds.
No more choosing algorithms and building complex pipelines. Our technology enables both experts and enthusiasts to focus entirely on their research objectives without being side-tracked by the technology. The BioStrand platform provides you with a range of intuitive, powerful, versatile, and multidimensional tools that allow you to define the scope, focus, and pace of your research without being restricted by any technological limitations. Parse, slice, dice, sort, filter, drill down, pan out, and do whatever it takes to define and pursue the research pathways that you think have the maximum potential for a breakthrough.
Leverage the power of the BioStrand platform’s state-of-the-art AI tools to quickly and intuitively synthesise knowledge from a multitude of data sources and across structured and unstructured data types.
With BioStrand research, researchers and bioinformaticians finally have access to a user-centric, multidimensional, secure, end-to-end data-to-insight research platform that enables a personalised and productive research experience by leveraging the power of modern digital technologies in the background
Harnessing the potential of population genomics
Population genomic data will continue to grow as more and more countries, especially in the developing world, realise the positive impact large-scale sequencing can have on genomics research, personalised patient care and public precision health. However, data science is key to realising the inherent value of genomic data at scale. Conventional approaches to genomic research and analysis are severely limited in terms of their ability to efficiently extract value from genomics big data. And research is often hampered by the need for highly skilled human capital that is hard to come by.
With the BioStrand platform, genomics research finally has an integrated solution that incorporates all research-related workflows, unifies discrete data sources and provides all the tools, features and functionality required for researchers to focus on what really matters – pushing the boundaries of genomics research, personalised patient care, and public precision health.
Subscribe to our blog:






