Powering biotherapeutic intelligence
Stories from the leading edge
Blogs
- All
- AI
- Omics analysis
- ML
- Drug discovery
- NLP
- Precision medicine
- Bioinformatics
- Life sciences data management
- In silico
- BioStrand Company
- Company culture
- Employee spotlight
- R&D
- Knowledge Graphs
- Large language models
- Antibody discovery
- Microbiome
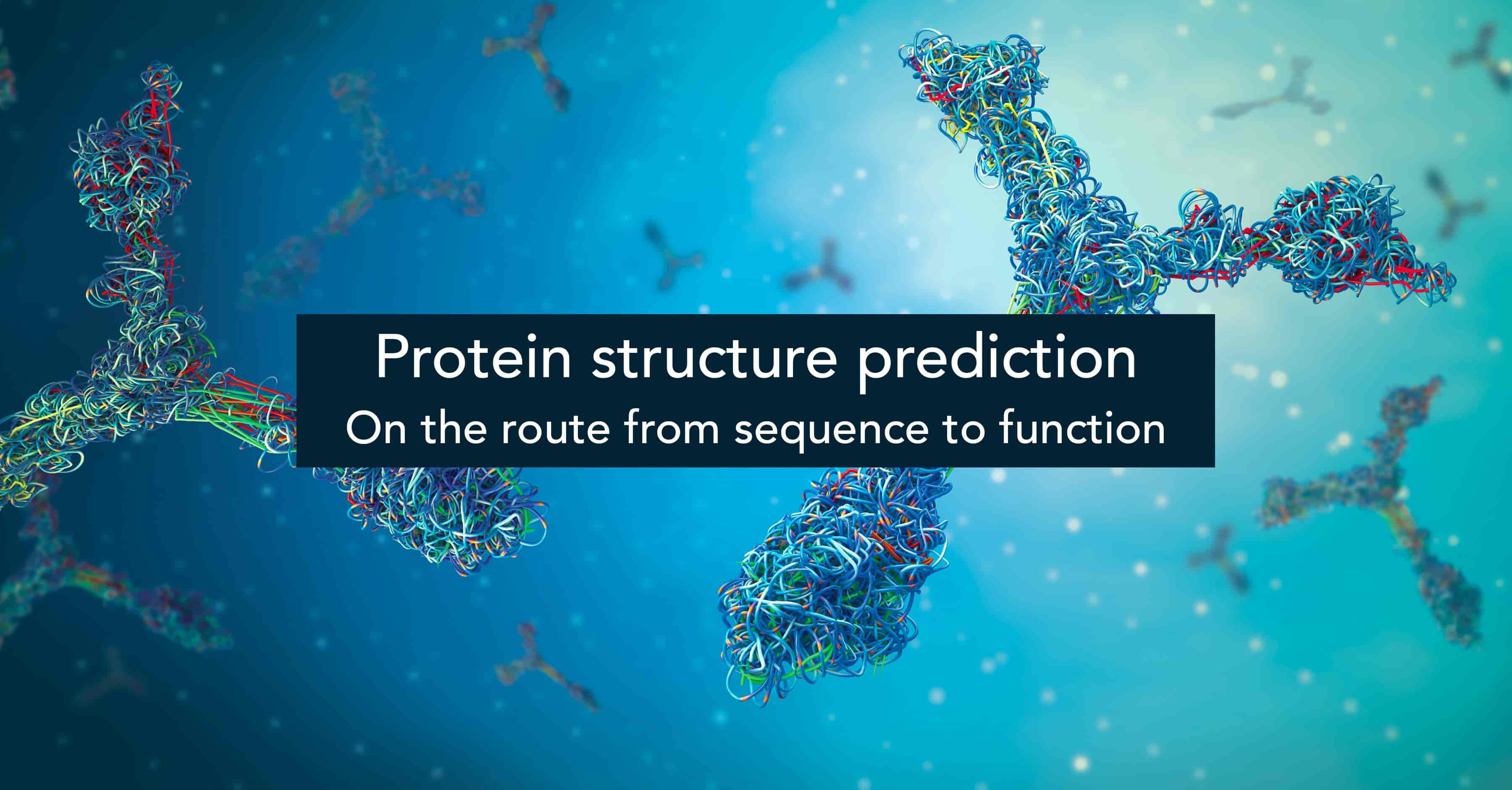
- Protein structure prediction
- Retrieval Augmented Generation
- AlphaFold
- Big data
- Biomarker
- Cloud computing
- Metadata
- Anaconda
- Biobank
- Biomarkers
- Conda
- Conda environment
- Data science
- De novo design
- Generative AI
- Multimodality
- Named entity recognition
- Ontologies
- Python package manager
- Relation extraction
- Semantics
Insights into genetics and genomics research, and their leading role in the rapid development of the field.
Popular
Subscribe to our blog: